Life as we know it can be distilled into just four classes of biomolecules: lipids, nucleic acids, proteins, and carbohydrates. Among these, sugars are the least studied, but the next frontier of biology will come from uncovering their functions in health and disease.
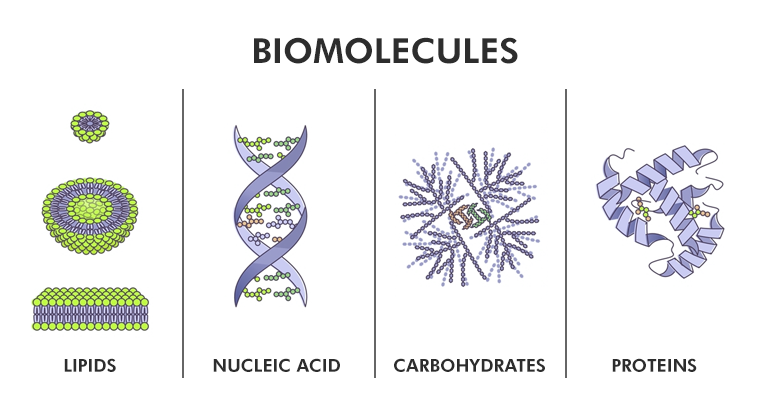
If DNA in a cell represents the blueprint of a building, and proteins + lipids make up its architecture, the sugars are the day-to-day workers in that building. They come and go, change their attitude and appearance often, and fundamentally power the whole assembly. Sugar studies, or glycobiology, are the important next step for life sciences. – sugars power and control all aspects of a cell! When they go wrong, diseases like cancer, diabetes, and neurodegeneration arise. We seek to understand sugars in order to treat and prevent diseases that stem from uncontrolled sugar use.
Chemical tools enable the study of complex “macro” systems with molecular precision. Coupled with advances in analytical methods, live-cell techniques, and data informatics, our generation of scientists has more power than ever to determine subtle effects in biology. Protein post-translational modifications (PTMs)–which often add/change only a few atoms of a kilodalton sized-system–are the cell’s way of actively fine-tuning signaling and gene expression in response to environmental cues.
One of the most common pathways to go awry in disease is metabolic regulation. Cancer cells become more metabolically active than they should be (a survival advantage), neurons experiencing tauopathic stress are less active than they should be (neurodegeneration), and adipocytes with misregulated sugar levels become resistant to insulin (type 2 diabetes). Facets of these diseases can be traced back to altered signaling pathways controlled by sugar levels in cells, which in turn modify proteins via PTMs.
By focusing on the development of chemical tools to “tag” sugar PTMs in living cells, we hope to understand these processes in molecular detail. Proteomics allows a view of which proteins are important for signaling, transcriptomics reveals how metabolic genes are switched on/off, and machine learning/data informatics strategies allow us to correlate these two very different types of “big” datasets and mine them for therapeutic action. Our glycopeptidomimetic design platform allows us to subsequently target these pathways in disease model systems.
Mentoring Statement:
The Fehl Lab mentoring goal is to craft independent and creative chemical biologists. Our core values are: clear communication, fostering success through student-driven research plans, a supportive environment, and a strongly inclusive and healthy lab culture.
Group members can expect kind but critical feedback about experimental design and results in order to inspire growth. At the end of the day, what you put in determines what you get out. To this end, trainees set their Action Plans regularly to make sure career goals are met. Weekly group meetings and biweekly one-on-one meetings with the PI enable mentored direction as well as freedom to explore and grow creatively.
Support for all career pathways are important for developing independent chemical biologists, which spans diverse disciplines. We work on networking, professional development, and scientific output in terms of papers, patents, and presentations. Students, and the PI, support each other’s learning through weekly “Data Camp” meetings to critically evaluate the literature, solve practice problems, and critique their own data. The PI and experienced members (advanced grad students, postdocs) advocate for the success of all trainees in the Fehl Group.
We also strongly value a diverse and inclusive group culture. Weekly “D&I Minutes” kick off our group meetings, and we hold regular events in the Wayne State and Detroit communities to give back with outreach and social justice conversations. Trainees can expect to become members of an aware, healthy, and constantly questioning community.
Welcome to the Fehl Lab!